-
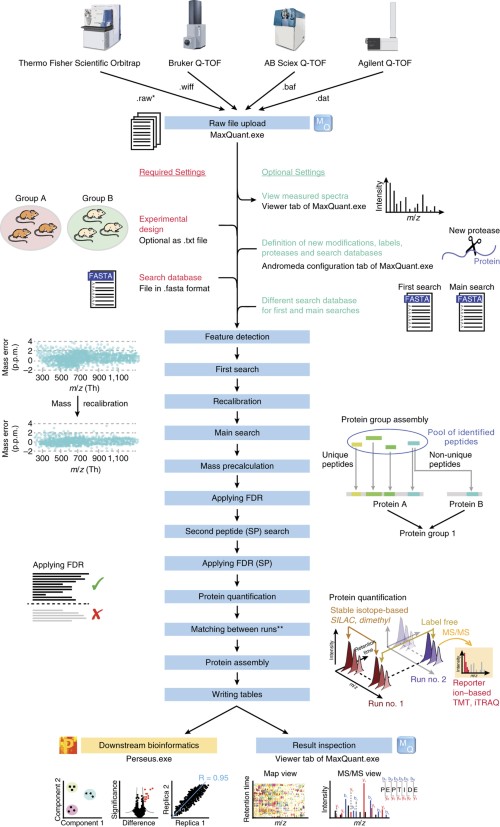 Mass Spectrometry Protocols and Methods Springer Nature Experiments
Mass Spectrometry Protocols and Methods Springer Nature Experiments
ID: MijwHFYNIi
From: experiments.springernature.com
-
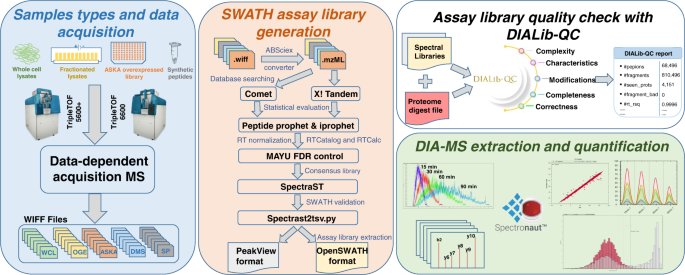 comprehensive spectral assay library to the Escherichia coli proteome by DIA/SWATH-MS | Scientific Data
comprehensive spectral assay library to the Escherichia coli proteome by DIA/SWATH-MS | Scientific Data
ID: ekzyErSXy5
From: nature.com
-
 Peptide-Centric Proteome Analysis: Alternative Strategy the Analysis of Tandem Mass Spectrometry Data* - Molecular & Cellular Proteomics
Peptide-Centric Proteome Analysis: Alternative Strategy the Analysis of Tandem Mass Spectrometry Data* - Molecular & Cellular Proteomics
ID: VTLjxdxv0q
From: mcponline.org
-
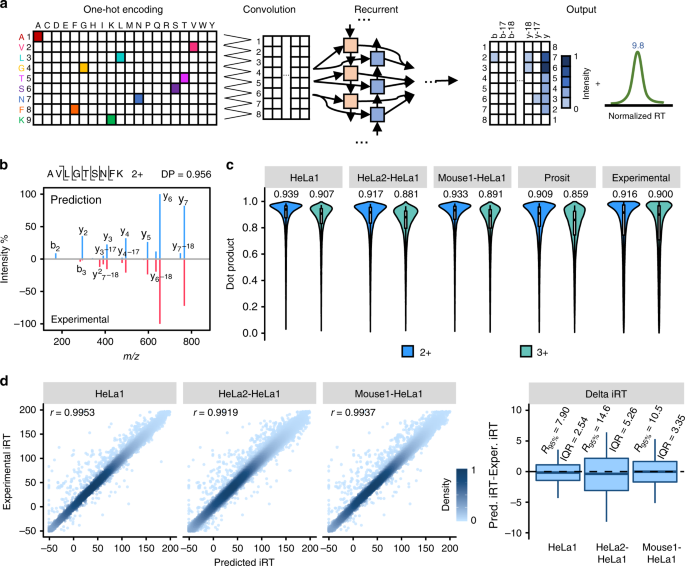 In silico spectral by deep learning facilitate data-independent acquisition proteomics | Nature Communications
In silico spectral by deep learning facilitate data-independent acquisition proteomics | Nature Communications
ID: SQdojb2rz4
From: nature.com
-
 Data-Independent Acquisition: Superior in Mass Spectrometry? | Technology
Data-Independent Acquisition: Superior in Mass Spectrometry? | Technology
ID: nE3A3KA4Kk
From: technologynetworks.com
-
 DIAproteomics: A multi-functional analysis pipeline data-independent-acquisition proteomics and peptidomics | bioRxiv
DIAproteomics: A multi-functional analysis pipeline data-independent-acquisition proteomics and peptidomics | bioRxiv
ID: BiiNqpvCj3
From: biorxiv.org
-
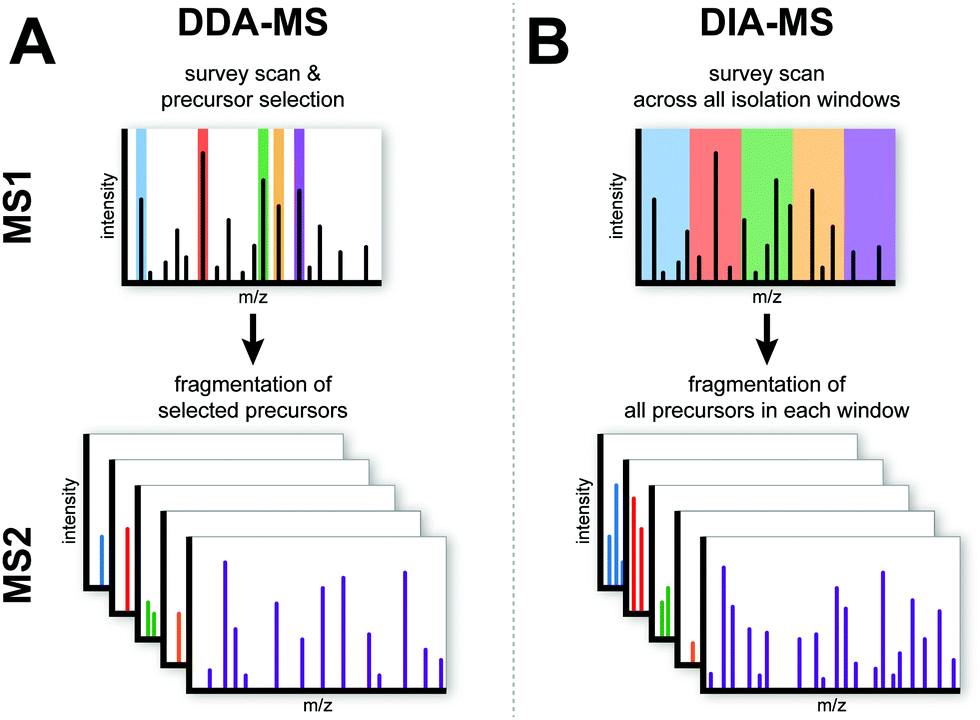 Data-independent acquisition mass spectrometry (DIA-MS) proteomic applications in oncology - Molecular (RSC Publishing)
Data-independent acquisition mass spectrometry (DIA-MS) proteomic applications in oncology - Molecular (RSC Publishing)
ID: jo61awW9Dx
From: pubs.rsc.org
-
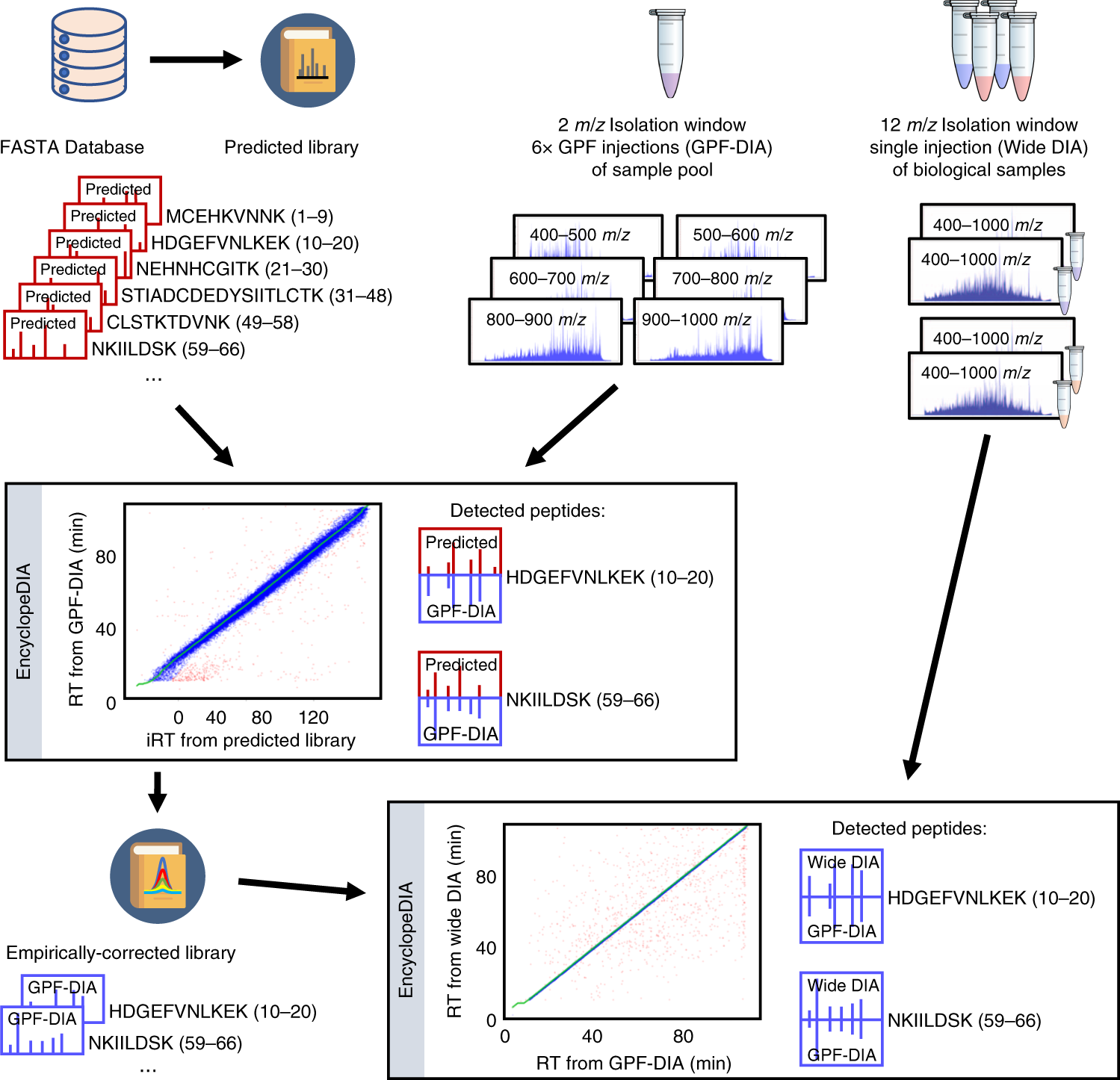 Generating for DIA MS with empirically corrected peptide | Nature Communications
Generating for DIA MS with empirically corrected peptide | Nature Communications
ID: IUGnwyS1wf
From: nature.com
-
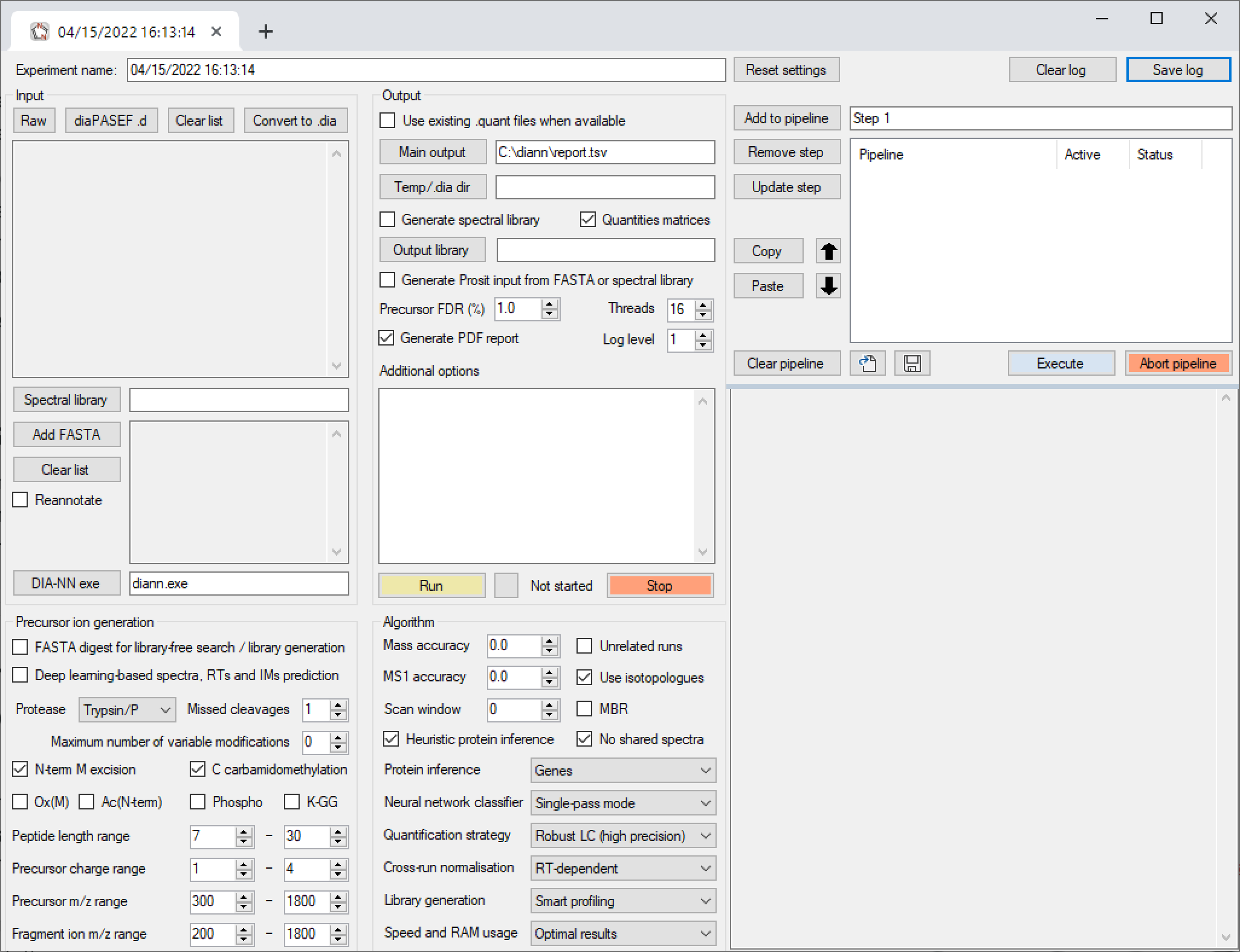 GitHub - vdemichev/DiaNN: DIA-NN - a universal automated software for DIA proteomics data analysis.
GitHub - vdemichev/DiaNN: DIA-NN - a universal automated software for DIA proteomics data analysis.
ID: tRhn8MtrW4
From: github.com
-
 Data‐independent acquisition‐based for quantitative proteomics: a | Molecular Systems Biology
Data‐independent acquisition‐based for quantitative proteomics: a | Molecular Systems Biology
ID: 0SkdGpCyOe
From: embopress.org
-
 Ultra-High-Throughput Clinical Reveals Classifiers of COVID-19 Infection - ScienceDirect
Ultra-High-Throughput Clinical Reveals Classifiers of COVID-19 Infection - ScienceDirect
ID: x9qIxsIotA
From: sciencedirect.com
-
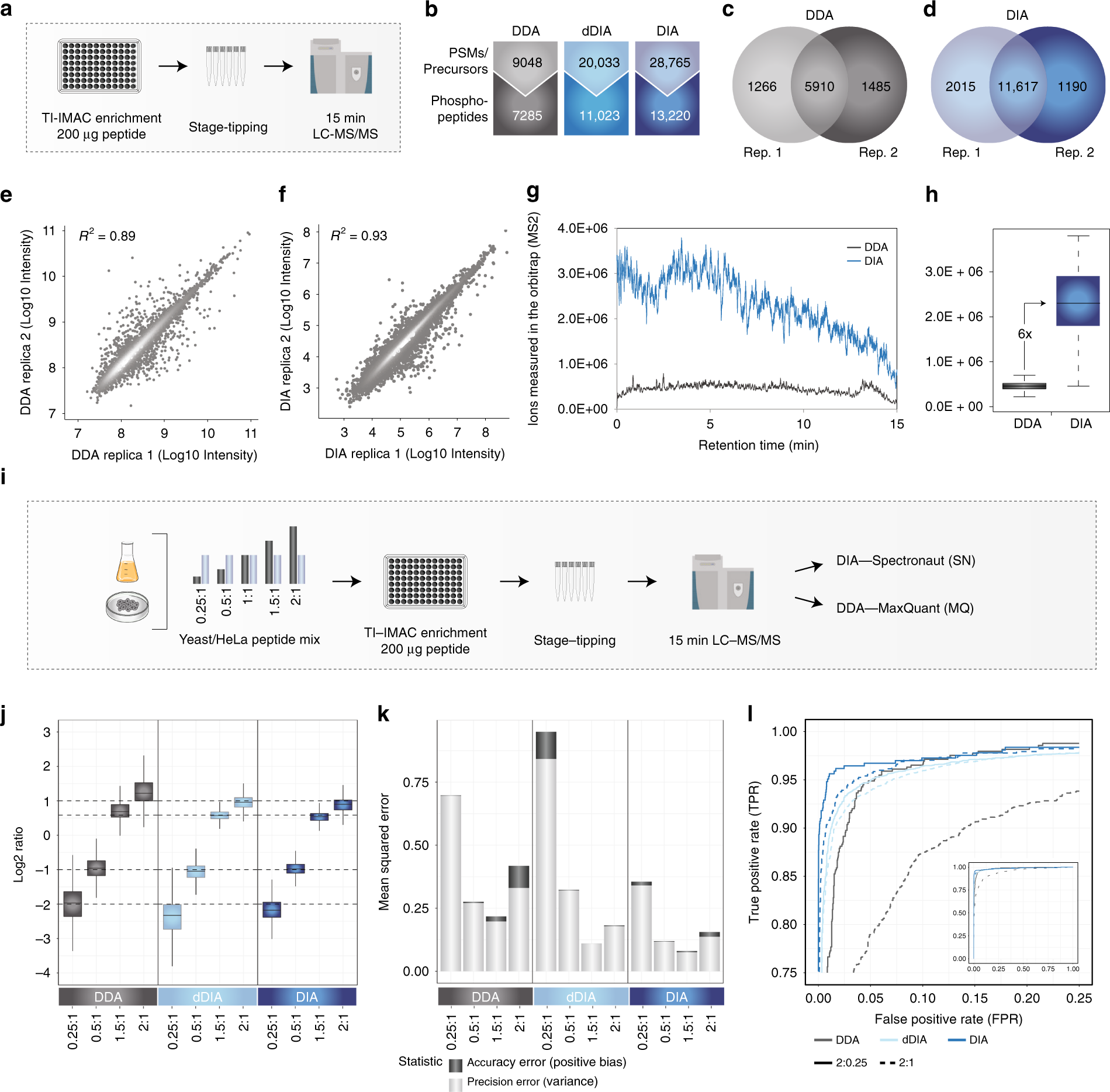 Rapid and site-specific deep phosphoproteome profiling by data-independent acquisition without the need for spectral |
Rapid and site-specific deep phosphoproteome profiling by data-independent acquisition without the need for spectral |
ID: I43gCZgtgc
From: nature.com
-
 Library Combining DIA-MS Data a Targeted Library Substantially Deepens the Proteome Coverage - ScienceDirect
Library Combining DIA-MS Data a Targeted Library Substantially Deepens the Proteome Coverage - ScienceDirect
ID: yW2OcVyCYd
From: sciencedirect.com
-
 SWATH analysis and spectrometry - is it?
SWATH analysis and spectrometry - is it?
ID: i00bIyX5kC
From: alphalyse.com
-
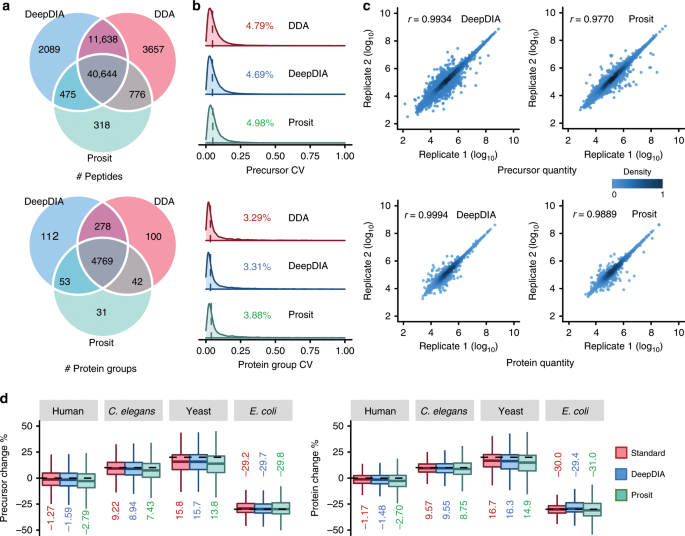 In silico spectral by deep learning facilitate data-independent acquisition proteomics | Nature Communications
In silico spectral by deep learning facilitate data-independent acquisition proteomics | Nature Communications
ID: xErpOsUDfG
From: nature.com
-
 Recap: DIA
Recap: DIA
ID: KrpKT3QCtu
From: inf.fu-berlin.de
-
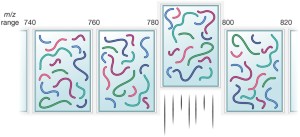 mass spectrometry | Nature Methods
mass spectrometry | Nature Methods
ID: 0iKcqchB3U
From: nature.com
-
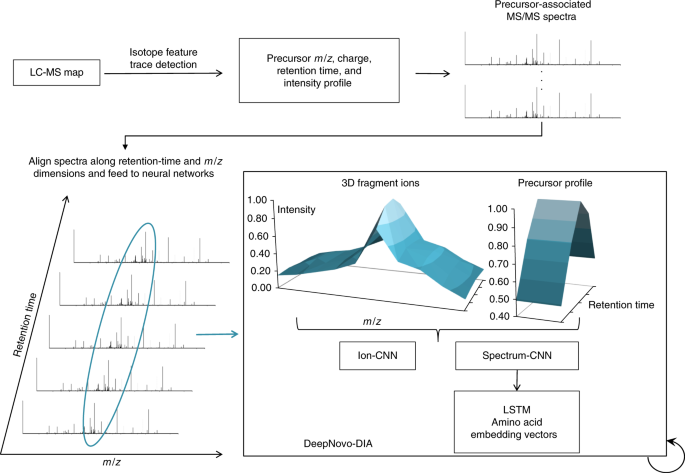 Deep enables de novo peptide sequencing from data-independent-acquisition spectrometry Nature Methods
Deep enables de novo peptide sequencing from data-independent-acquisition spectrometry Nature Methods
ID: TCX5SSLJ73
From: nature.com
-
 Mass spectrometry‐based interaction networks for the study human diseases | Molecular Systems Biology
Mass spectrometry‐based interaction networks for the study human diseases | Molecular Systems Biology
ID: z4o5pXvnFP
From: embopress.org
-
 PDF) Multiplexed MS/MS for Data Independent Acquisition
PDF) Multiplexed MS/MS for Data Independent Acquisition
ID: cspoAqIR97
From: researchgate.net
-
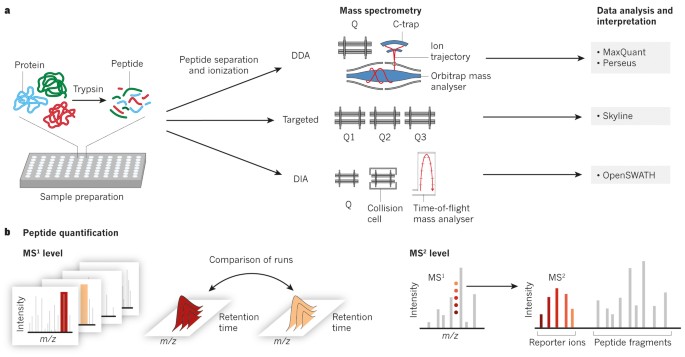 Mass-spectrometric exploration of proteome and | Nature
Mass-spectrometric exploration of proteome and | Nature
ID: ET5GQJVHu3
From: nature.com
-
 Data‐independent acquisition‐based for quantitative proteomics: a | Molecular Systems Biology
Data‐independent acquisition‐based for quantitative proteomics: a | Molecular Systems Biology
ID: k3vjZ4U3Vl
From: embopress.org
-
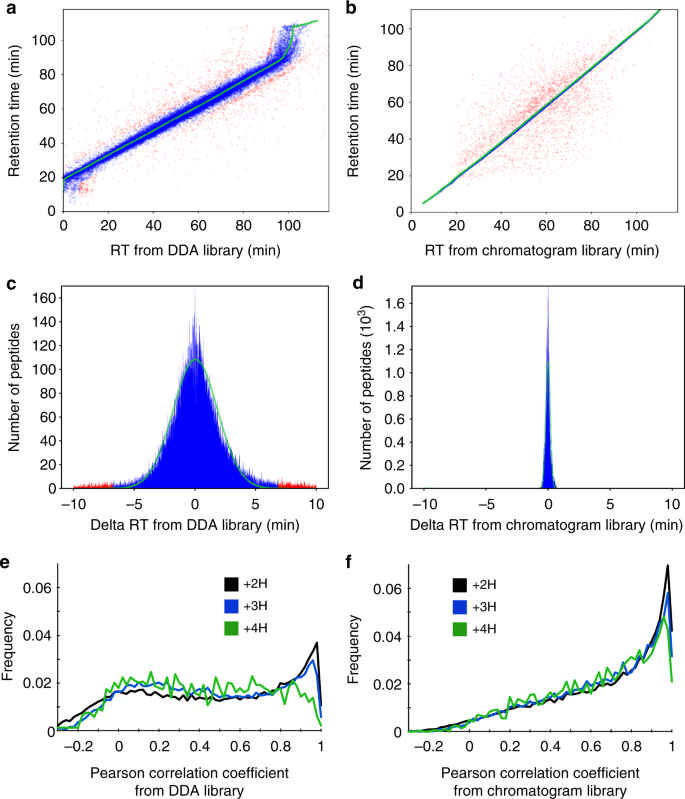 Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry | Communications
Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry | Communications
ID: TBeUZJjN1n
From: nature.com
-
 Quantifying Plant Dynamic Proteomes by SWATH-based Mass Spectrometry: Plant Science
Quantifying Plant Dynamic Proteomes by SWATH-based Mass Spectrometry: Plant Science
ID: O03jy1ogiW
From: cell.com
-
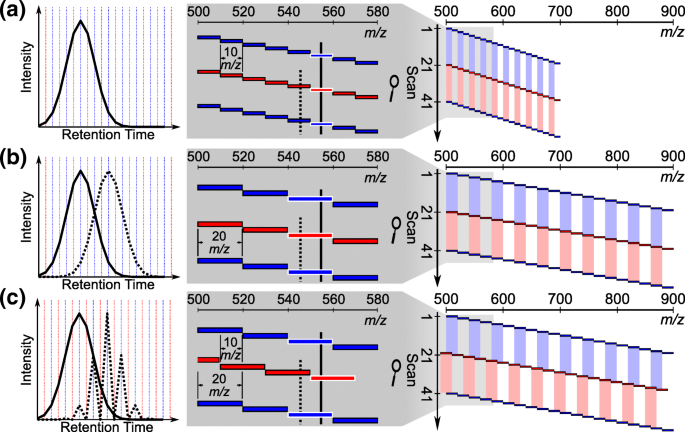 Improving Precursor Selectivity Data-Independent Overlapping Windows | SpringerLink
Improving Precursor Selectivity Data-Independent Overlapping Windows | SpringerLink
ID: nSOWIvSr1M
From: link.springer.com